Hepatocyte Isolation and Purification – A Worldwide Research Priority
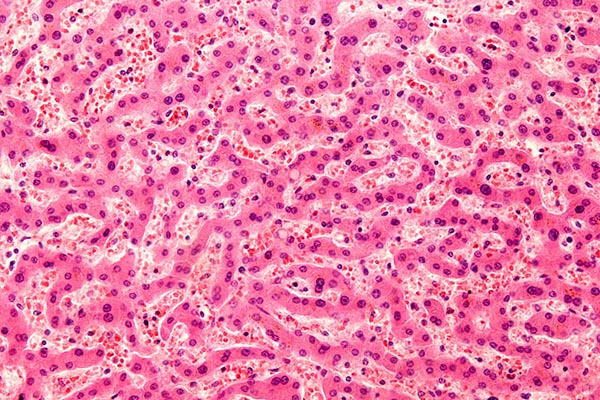
The largest solid organ in the body is the liver. Made up primarily (i.e., at least 80%) of hepatocytes of various types, the liver is the site of hepatocellular carcinoma (HCC), the fourth ranked cause of cancer deaths worldwide. A large and growing HCC caseload has led to a vast World-wide research effort to better understand hepatocyte structure, function and pathology. Hepatocyte research is also driven by another major factor, namely the need to better manage preclinical testing of pharmaceutical compounds for hepatotoxicity, a major reason for retractions of marketed drugs not to mention the outright failure of drugs during clinical trials. Both oncology and drug testing would benefit if the culturing of human hepatocytes could be perfected. Finally, cultured human hepatocytes hold the promise of development of bioengineered livers and use in cell transplantation in diseased livers for correction of metabolic disorders or as a bridge to organ transplantation.
Experiments in the late 1960’s resulted in the first successful hepatocyte isolation protocols and this laboratory work culminated around 1976 when Seglen published the two-step perfusion protocol that is the basis for the hepatocyte isolation protocols used today. However, even with improved protocols, and newer techniques to culture isolated primary hepatocytes, the result is often loss of phenotype and altered gene expression, problems unresolved to this day.
Better isolation/purification methods must be invented. Despite decades of research, it is still not possible to maintain the maturity of hepatocytes in long-term culture. During culture, primary hepatocytes undergo a process called dedifferentiation where they rapidly lose expression of a multitude of genes, including those that are involved in key functions such as xenobiotic metabolism, glucose metabolism, urea synthesis, and amino acid metabolism.
Current research suggests that the loss of phenotype for cultured hepatocytes occurs via two distinct processes: 1) an inflammatory response centered around NF-κβ (nuclear factor kappa-light-chain-enhancer of activated B cells) activation. This is induced by ischemia/reperfusion damage, hypoxia, and tissue disruption during the isolation process, resulting in the formation of reactive compounds such as reactive oxygen species (ROS), which can directly lead to injury in hepatocytes or induce the secretions of pro-inflammatory cytokines from the liver cells, especially Kupffer cells. Notably, this inflammatory response is often worsened by the contamination of the collagenase used during perfusion by bacterial lipopolysaccharide activating Kupfer cells and inducing the release of pro-inflammatory cytokines; and 2) Loss of phenotype occurs during a proliferative phase that is based on activation of mitogen-activated protein kinase (MAPK) signaling among other pathways. A result is massive changes involving mRNA, proteins, and noncoding RNAs driving the hepatocytes away from their normally highly differentiated state toward a proliferative state concurrently downregulating genes responsible for metabolic processes associated with mature liver functions such as cytochrome P450s, UGTs, SULTs, and enzymes involved in glucose metabolism. A sub-optimal extracellular matrix can also contribute to the dedifferentiation of mature hepatocytes. For example, a stiff extracellular matrix can lead to the formation of stress fibers and tension in the cells.
Many methods have been investigated to extend the mature hepatic phenotype in culture, including media additives and novel culturing techniques. Media additives are classified as physiological versus non-physiological. The physiological additives include insulin, growth factors (e.g., EGF and HGF), glucocorticoids (especially dexamethasone), glutamine, selenium and transferrin, and fetal bovine serum. The nonphysiological additives include dimethyl sulfoxide (DMSO) and enzyme inducers such as rifampicin and phenobarbital.
Liver ECM is also a Principal Research Focus
Concurrent with research to culture hepatocytes, perfecting a liver extracellular matrix (ECM) is also a goal. A biomimetic ECM is necessary to simulate healthy versus damaged liver for liver disease research, drug toxicity studies and cancer metastasis modelling. Currently used liver ECM hydrogels are based on time-consuming thermal gelation and that limits the control of mechanical properties. A new technique though (see Ravichandran A, Murekatete B, Moedder D, Meinert C, Bray LJ. Photocrosslinkable liver extracellular matrix hydrogels for the generation of 3D liver microenvironment models. Sci Rep. 2021 Jul 30;11(1):15566. doi: 10.1038/s41598-021-94990-z. PMID: 34330947; PMCID: PMC8324893) demonstrates use of detergent-based protocols to produce a decellularized porcine liver ECM, which in turn can be solubilized and functionalized with methacrylic anhydride to generate photocrosslinkable methacrylated liver ECM (LivMA) hydrogels. The research demonstrated the efficacy of two protocols to decellularize porcine liver tissue using varying combinations of commonly used chemical agents such as Triton X-100, Sodium Dodecyl Sulphate (SDS) and Ammonium hydroxide. A successful formation of stable, reproducible LivMA hydrogels was demonstrated via both protocols by photocrosslinking. The LivMA hydrogels obtained from the two decellularization protocols showed distinct mechanical properties. The compressive modulus of the hydrogels was directly dependent on the hydrogel concentration, thereby demonstrating the tunability of mechanical properties of these hydrogels. Immortalized Human Hepatocytes were encapsulated in the LivMA hydrogels and cytocompatibility of the hydrogels was demonstrated after one week of culture. In summary, the LivMA hydrogel system demonstrated in this research provides a simple, photocrosslinkable platform, which can potentially be used to simulate healthy versus damaged liver for liver disease research, drug studies and cancer metastasis modelling.